Introduction
The experiment configuration system provides automated generation and management of configuration files for the PhenoCycler data analysis pipeline. This system enables researchers to efficiently manage multiple experiments through CSV-based design tables and YAML templates.
Experiment Configuration Architecture
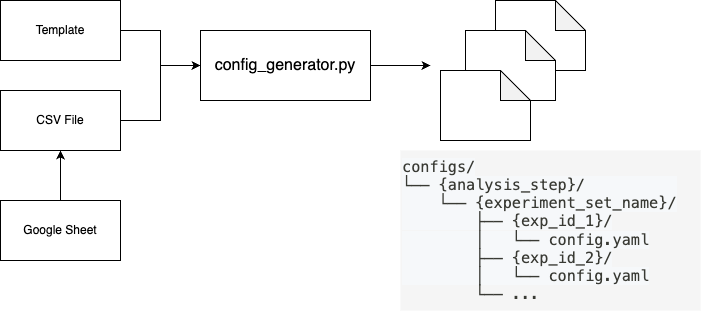
Key Features
The configuration generator (config_generator.py) provides:
- Batch Configuration Generation: Generate multiple experiment configurations from CSV experiment design tables and YAML templates
- Automatic Type Conversion: Handle parameter conversion for different data types automatically
- Nested Configuration Support: Support multi-level configuration structures using
::separators - Batch Script Generation: Generate experiment list strings for batch processing
Directory Structure
The experiment configuration system is located in the exps/ directory:
exps/
├── README.md # Documentation
├── config_generator.py # Main configuration generator script
├── config_generators/ # Specialized configuration generator modules
│ ├── config_generator_main.py
│ ├── config_generator_preprocess.py
│ └── generator_utils.py
├── csvs/ # Experiment design tables exported from Google Sheet
│ ├── main_ft.csv
│ ├── preprocess_ft.csv
│ ├── test_analysis.csv
│ └── ...
├── configs/ # Generated configuration files
│ ├── analysis/
│ ├── main/
│ └── preprocess/
├── *_template.yaml # Configuration template files
│ ├── main_template.yaml
│ ├── preprocess_template.yaml
│ └── analysis_template.yaml
└── archive/ # Historical configurations and experiments
Workflow Overview
The typical workflow for experiment configuration involves:
- Design Experiments: Create experiment designs in the Google Sheet
- Export CSV: Export the design table as CSV files to the
csvs/directory - Configure Generator: Set the experiment parameters in
config_generator.py - Generate Configs: Run the generator to create YAML configuration files
- Run Pipeline: Use the generated configurations with the Aegle pipeline
Pipeline Integration
The experiment configuration system integrates with three major components of the Aegle Pipeline:
- Preprocessing (
preprocess_X): Tissue extraction and basic image processing - Main Analysis (
main_X): Image segmentation, patch extraction, and cell profiling - Analysis (
analysis_X): Downstream statistical analysis and visualization
Each component has its own design table format and configuration template, ensuring proper parameter handling across the entire pipeline.